Molecular Structure of Nucleic Acids: a Structure for Deoxyribose Nucleic Acid - J.d. Watson, F.h.c. Crick.
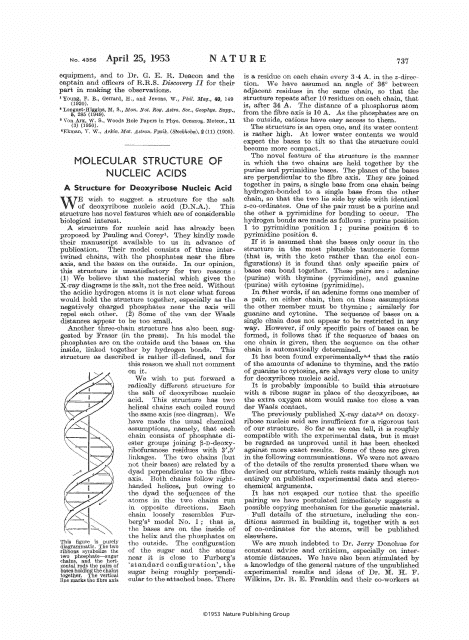
The document "Molecular Structure of Nucleic Acids: a Structure for Deoxyribose Nucleic Acid - J.D. Watson, F.H.C. Crick." describes the ground-breaking discovery of the molecular structure of DNA (deoxyribose nucleic acid) by James D. Watson and Francis H.C. Crick. This discovery, published in 1953, provided the first detailed understanding of how DNA is structured and how it carries genetic information. The document outlines the famous double helix structure of DNA, which consists of two strands twisted together in a spiral staircase-like formation. This groundbreaking work laid the foundation for our understanding of genetics and revolutionized the field of molecular biology.
The molecular structure of nucleic acids, specifically the structure for deoxyribose nucleic acid (DNA), was filed by James D. Watson and Francis H.C. Crick. They are credited with discovering and proposing the double helix structure of DNA in 1953. This breakthrough laid the foundation for modern genetics and our understanding of heredity.
FAQ
Q: What is the book 'Molecular Structure of Nucleic Acids' about?
A: The book 'Molecular Structure of Nucleic Acids' is about the discovery of the structure of deoxyribose nucleic acid (DNA) by James Watson and Francis Crick.
Q: Who are the authors of the book 'Molecular Structure of Nucleic Acids'?
A: The authors of the book 'Molecular Structure of Nucleic Acids' are James D. Watson and Francis H.C. Crick.
Q: What is the significance of the book 'Molecular Structure of Nucleic Acids'?
A: The book 'Molecular Structure of Nucleic Acids' is significant because it outlines the double-helix structure of DNA, which is the fundamental basis of genetic information and the understanding of heredity.
Q: When was the book 'Molecular Structure of Nucleic Acids' published?
A: The book 'Molecular Structure of Nucleic Acids' was published in 1953.
Q: What is the full title of the book 'Molecular Structure of Nucleic Acids'?
A: The full title of the book is 'Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid'.